The topology of Chloroplast RNA metabolism
Chloroplast gene expression takes place in nucloids (the DNA compartment), in the stroma (the souble compartment) and on thalykoid membranes. We have analyzed the association of RNAs with the transcriptionally active chromosome, which is a preparation representing actively transcribing nucleoids (Lehniger et al. 2017). Also, we identified RNAs associated with the main RNA polymerase, the PEP (Finster et al. 2013). These analyses demonstrated that nascent RNAs can be separated from mature RNAs via their association with the nucleoid compartment. We also investigated the partitioning of RNAs between the membrane and soluble fraction of the chloroplast, which indicated that different RNAs processed from polycistronic precursors show transcript-autonomous distribution between stroma and membrane fractions (Legen and Schmitz-Linneweber 2017).
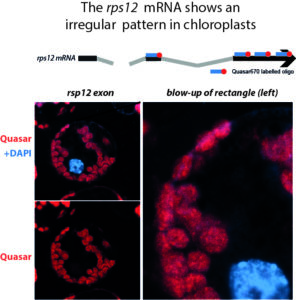
These analyses already made a case for a non-random distribution of RNA processing factors. Dedicated RNA compartments, i.e. RNA granules have been described in Chlamydomonas reinhardtii, but are so far not found in chloroplasts of land plants. We seek to identify the suborganellar localization of RNAs and their protein ligands using superresolution microscopy. First analyses of RNA localization using single molecile FISH already point to interesting structures in RNA metabolism.
Publications
Legen J, Schmitz-Linneweber C
Stable Membrane-Association of mRNAs in Etiolated, Greening and Mature Plastids
In: Int J Mol Sci. 2017 Aug 31;18(9). pii: E1881. doi: 10.3390/ijms18091881.
Lehniger MK, Finster S, Melonek J, Oetke S, Krupinska K, Schmitz-Linneweber C
Global RNA association with the transcriptionally active chromosome of chloroplasts
In: Plant Mol Biol. 2017 Sep 8. doi: 10.1007/s11103-017-0649-x
Finster S, Eggert E, Zoschke R, Weihe A, Schmitz-Linneweber C
Light-dependent, plastome-wide association of the plastid-encoded RNA polymerase with chloroplast DNA
In: Plant J., 2013 Oct 7; doi: 10.1111/tpj.12339 [Epub 2013 Nov 8]