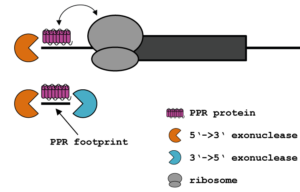
Plant PPR proteins
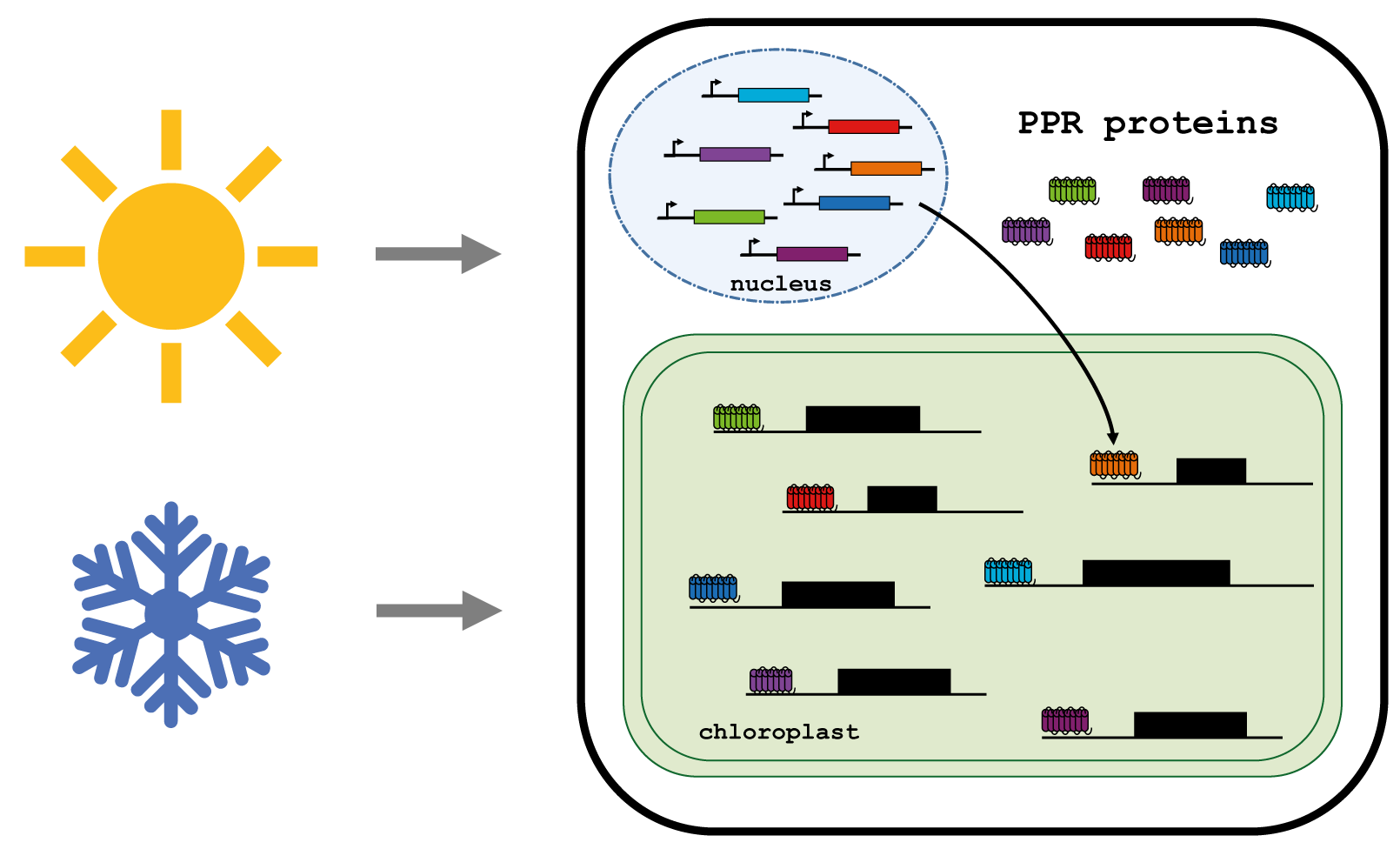
PPR proteins as regulators of the acclimation of chloroplast RNA metabolism
Pentatricopeptide repeat (PPR) proteins are RNA-binding proteins with high specificity. This protein family is present in almost every eukaryotic genome. Encoded in the nuclear genome, most members are imported post-translationally into the two endosymbiotic organelles, the mitochondrion and the chloroplast. In these DNA-containing organelles they bind RNA transcripts and influence organellar gene-expression post-transcriptionally.
Plants as sessile organisms need to acclimate to environmental conditions. In temperate regions, plants experience low temperatures and shorter day-length during winter and elevated temperatures with increased light intensities during summer. Temperature and light quantities are integrated by plants and result in acclimation responses that include reprogramming of gene expression patterns.
The genes encoded in the chloroplast genome are regulated predominantly on the post-transcriptional level. PPR proteins impact RNA stability and influence the rate of translation of individual chloroplast mRNAs. Our goal is to understand how PPR proteins reprogram chloroplast gene-expression in response to environmental signals, especially temperature.
Identification of PPR protein binding-sites
Exact binding-sites of PPR proteins can be identified by investigating in vivo footprints of these RNA-binding proteins. The footprints result from the intrinsic property of many PPR proteins to block progression of exonucleases, a mean to stabilize chloroplast mRNAs (figure above). Sequencing small RNAs allows the identification of potential PPR protein target sites on a genome-wide level. The figure below shows small RNA accumulation in a typical chloroplast operon. Clusters of small RNAs predict binding sites for PPR and related RNA-binding proteins (marked with arrows).
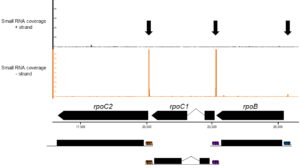
Click here to access the interactive genome browser of chloroplast and mitochondrial small RNAs – including many PPR footprints – described in our 2016 NAR paper.
Publications:
Click here for a list of publications from the group.